Model info
| Transcription factor | CEBPB | ||||||||
| Model | CEBPB_HUMAN.H10DI.A | ||||||||
| Model type | Dinucleotide PWM | ||||||||
| LOGO | 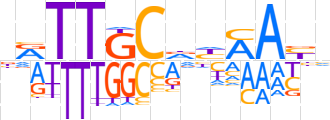 | ||||||||
| LOGO (reverse complement) | 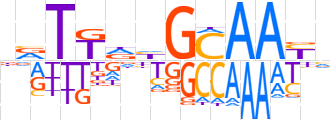 | ||||||||
| Origin | ChIP-Seq | ||||||||
| Model length | 12 | ||||||||
| Quality | A | ||||||||
| Consensus | nRTTGCdYAAYh | ||||||||
| wAUC | 0.9425690387646967 | ||||||||
| Best AUC | 0.9656326602270917 | ||||||||
| Benchmark datasets | 19 | ||||||||
| Aligned words | 516 | ||||||||
| TF family | C/EBP-related{1.1.8} | ||||||||
| TF subfamily | C/EBP{1.1.8.1} | ||||||||
| HGNC | 1834 | ||||||||
| EntrezGene | 1051 | ||||||||
| UniProt ID | CEBPB_HUMAN | ||||||||
| UniProt AC | P17676 | ||||||||
| Comment | |||||||||
| Downloads | pcm
pwm alignment threshold to P-value grid | ||||||||
| Standard thresholds |
|
PCM
| AA | AC | AG | AT | CA | CC | CG | CT | GA | GC | GG | GT | TA | TC | TG | TT | |
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| 01 | 45.394 | 8.518 | 43.268 | 4.968 | 62.26 | 12.281 | 10.171 | 0.0 | 99.96 | 6.591 | 43.809 | 9.696 | 83.162 | 18.044 | 50.814 | 1.064 |
| 02 | 0.0 | 0.0 | 3.665 | 287.111 | 0.0 | 0.0 | 0.0 | 45.434 | 0.0 | 0.0 | 0.838 | 147.224 | 0.0 | 0.0 | 0.0 | 15.727 |
| 03 | 0.0 | 0.0 | 0.0 | 0.0 | 0.0 | 0.0 | 0.0 | 0.0 | 0.0 | 0.0 | 0.0 | 4.503 | 0.0 | 3.829 | 0.908 | 490.76 |
| 04 | 0.0 | 0.0 | 0.0 | 0.0 | 0.0 | 0.0 | 2.94 | 0.89 | 0.0 | 0.0 | 0.908 | 0.0 | 36.546 | 1.092 | 397.982 | 59.642 |
| 05 | 1.845 | 34.702 | 0.0 | 0.0 | 0.0 | 1.092 | 0.0 | 0.0 | 0.0 | 394.415 | 1.898 | 5.518 | 0.0 | 57.923 | 0.0 | 2.609 |
| 06 | 0.0 | 0.944 | 0.0 | 0.901 | 255.222 | 47.845 | 117.541 | 67.523 | 1.898 | 0.0 | 0.0 | 0.0 | 4.505 | 1.79 | 0.935 | 0.897 |
| 07 | 51.074 | 113.178 | 4.828 | 92.545 | 13.085 | 20.986 | 0.0 | 16.508 | 25.973 | 40.711 | 0.0 | 51.791 | 5.769 | 36.858 | 0.865 | 25.828 |
| 08 | 41.664 | 52.553 | 0.0 | 1.685 | 190.516 | 21.218 | 0.0 | 0.0 | 2.906 | 2.787 | 0.0 | 0.0 | 82.344 | 103.351 | 0.977 | 0.0 |
| 09 | 311.546 | 2.077 | 2.937 | 0.869 | 177.255 | 1.725 | 0.0 | 0.928 | 0.977 | 0.0 | 0.0 | 0.0 | 1.685 | 0.0 | 0.0 | 0.0 |
| 10 | 6.745 | 183.15 | 84.278 | 217.29 | 0.0 | 2.077 | 0.0 | 1.725 | 0.0 | 0.0 | 1.0 | 1.938 | 0.0 | 0.0 | 0.0 | 1.798 |
| 11 | 2.89 | 2.903 | 0.953 | 0.0 | 68.902 | 58.275 | 5.553 | 52.497 | 27.19 | 31.27 | 16.176 | 10.641 | 56.43 | 83.743 | 34.247 | 48.33 |
PWM
| AA | AC | AG | AT | CA | CC | CG | CT | GA | GC | GG | GT | TA | TC | TG | TT | |
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| 01 | 0.37 | -1.268 | 0.322 | -1.776 | 0.683 | -0.915 | -1.097 | -4.4 | 1.154 | -1.511 | 0.334 | -1.143 | 0.971 | -0.54 | 0.481 | -3.081 |
| 02 | -4.4 | -4.4 | -2.055 | 2.207 | -4.4 | -4.4 | -4.4 | 0.37 | -4.4 | -4.4 | -3.251 | 1.54 | -4.4 | -4.4 | -4.4 | -0.675 |
| 03 | -4.4 | -4.4 | -4.4 | -4.4 | -4.4 | -4.4 | -4.4 | -4.4 | -4.4 | -4.4 | -4.4 | -1.867 | -4.4 | -2.015 | -3.195 | 2.742 |
| 04 | -4.4 | -4.4 | -4.4 | -4.4 | -4.4 | -4.4 | -2.252 | -3.209 | -4.4 | -4.4 | -3.195 | -4.4 | 0.155 | -3.062 | 2.533 | 0.64 |
| 05 | -2.651 | 0.104 | -4.4 | -4.4 | -4.4 | -3.062 | -4.4 | -4.4 | -4.4 | 2.524 | -2.628 | -1.678 | -4.4 | 0.611 | -4.4 | -2.357 |
| 06 | -4.4 | -3.167 | -4.4 | -3.2 | 2.089 | 0.422 | 1.316 | 0.764 | -2.628 | -4.4 | -4.4 | -4.4 | -1.866 | -2.676 | -3.174 | -3.203 |
| 07 | 0.486 | 1.278 | -1.803 | 1.078 | -0.854 | -0.392 | -4.4 | -0.627 | -0.182 | 0.262 | -4.4 | 0.5 | -1.637 | 0.163 | -3.228 | -0.188 |
| 08 | 0.285 | 0.515 | -4.4 | -2.725 | 1.797 | -0.381 | -4.4 | -4.4 | -2.262 | -2.299 | -4.4 | -4.4 | 0.961 | 1.188 | -3.143 | -4.4 |
| 09 | 2.288 | -2.552 | -2.253 | -3.225 | 1.725 | -2.706 | -4.4 | -3.179 | -3.143 | -4.4 | -4.4 | -4.4 | -2.725 | -4.4 | -4.4 | -4.4 |
| 10 | -1.49 | 1.758 | 0.984 | 1.929 | -4.4 | -2.552 | -4.4 | -2.706 | -4.4 | -4.4 | -3.127 | -2.61 | -4.4 | -4.4 | -4.4 | -2.672 |
| 11 | -2.267 | -2.263 | -3.161 | -4.4 | 0.784 | 0.617 | -1.672 | 0.514 | -0.137 | 0.001 | -0.647 | -1.054 | 0.585 | 0.978 | 0.091 | 0.432 |